Blogs and updates
Our blogs explain health conditions and clinical research in a way that is easy to follow, so readers can feel more informed
Common Endocrine Disorders and How to Treat Them
Hormones are like text messages between organs, allowing different parts of your body to communicate and stay in sync. They regulate your metabolism, mood, energy, sleep and even growth. They’re produced by a network of glands called the endocrine system and...
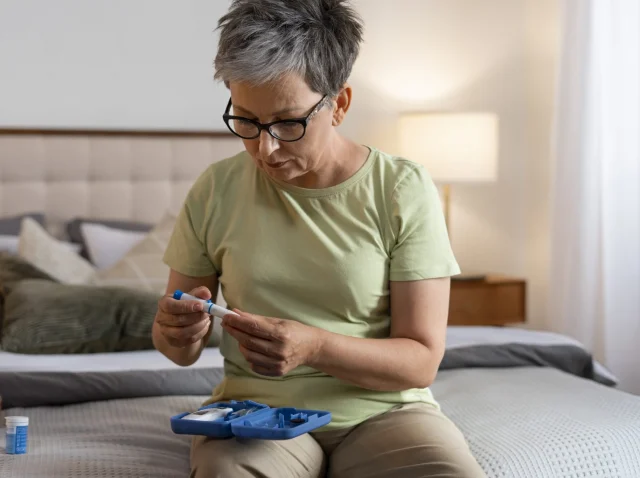
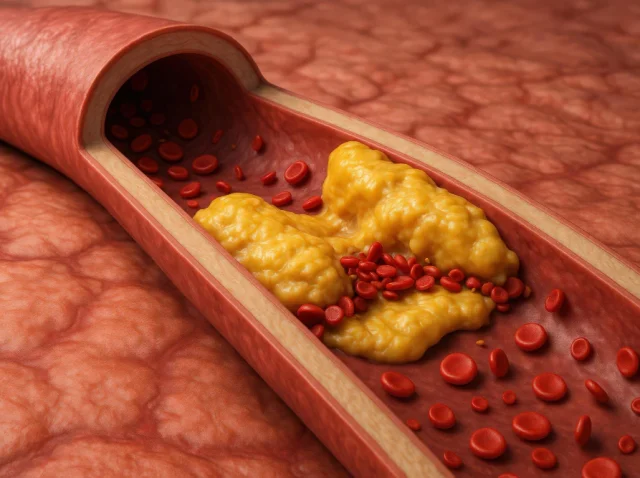
Why Regular Tests Matter for Detecting Hypertriglyceridemia Early
We view blood tests as routine, a standard part of annual checkups that we rarely question. But these simple tests can reveal so much about our health, often before symptoms appear. One of the most silent yet significant findings that can...
What Recent Research and Clinical Trials Teach Us About Thyroid Disease and Its Impact on Your Health
Imagine a busy city inside your body, filled with trillions of microscopic inhabitants working around the clock. This is the makeup of your gut microbiome, a complex community of microorganisms living in your digestive system. These tiny residents play a big...
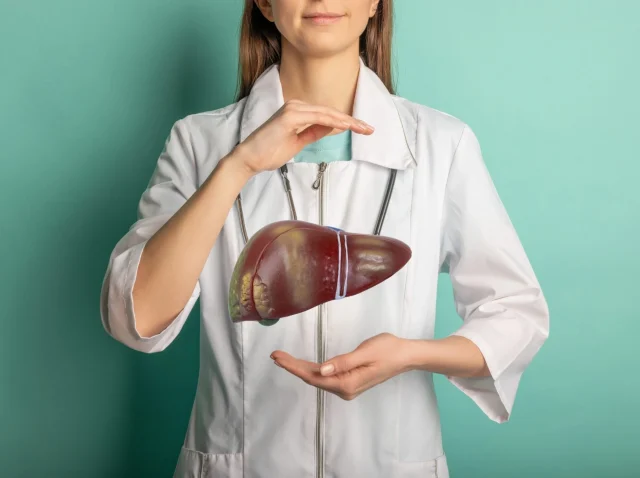
A Breakthrough in Liver Health Means New Hope for Liver Disease Clinical Trials and Treatment
When we think of vital organs, the liver might not be the first that comes to mind. But it actually does a lot of the heavy lifting when it comes to keeping us alive. The liver is responsible for processing nutrients,...
The Obstacles and Promise in Endocrine-Related Cancers
Cancer is a complicated disease and research is constantly changing our understanding of it. One of the latest is endocrine-related cancers (ERCs), cancers driven by hormones. Breast and prostate cancer have been hormone-driven for ages, but research is showing hormones may...
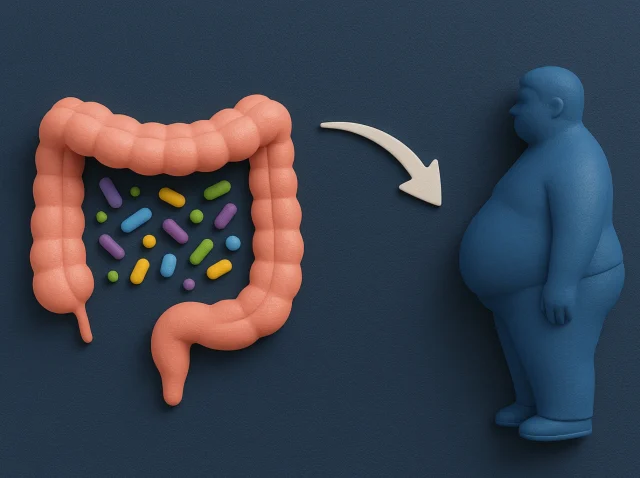
Your Gut, Your Health: How Microbes in Your Stomach Impact Metabolic Disease Risk
Your gut microbiome is an intricate community of microorganisms living in your digestive system. These microbes impact everything from digestion to immune function and even weight management. Let’s dive in and see how your gut microbiome affects metabolic diseases like obesity...
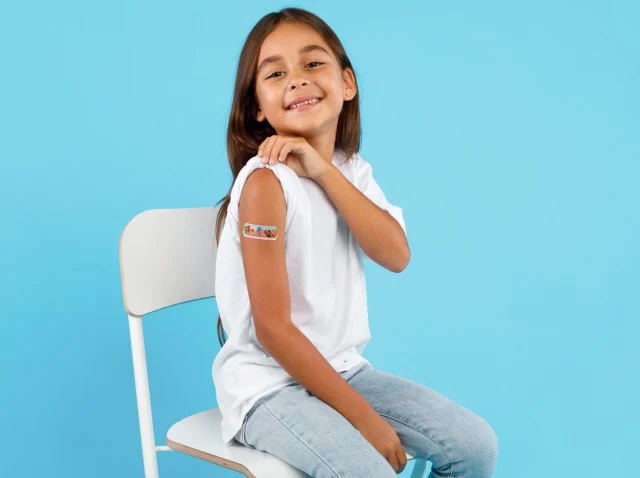
Why Children Need Dedicated Pediatric Vaccination Studies
Pediatric vaccination studies are the foundation of everything we know about keeping children safe from serious disease. From common illnesses like the flu to life-threatening conditions like polio, vaccines have saved an estimated 2 to 3 million lives worldwide every year....
Advancements in the Treatment of Endocrine Disorders
Endocrine Disorder Treatment: 5 Powerful Advances Changing Patient Care Endocrine disorder treatment is evolving rapidly as advances in technology, personalized medicine, and clinical research open new pathways for managing hormonal and metabolic conditions. The human body relies on the endocrine system...
What Hypertriglyceridemia Clinical Trials Are Teaching Us About Triglyceride-Lowering Medications
Hypertriglyceridemia Clinical Trials: 6 Proven Drug Insights Shaping Treatment Hypertriglyceridemia clinical trials are teaching researchers and clinicians more than ever about how to manage high triglyceride levels and reduce the serious health risks they carry. Hypertriglyceridemia occurs when triglyceride levels exceed...
How Precision Medicine and Obesity Trials are Changing Treatment Options
Obesity Clinical Trials: 5 Powerful Precision Medicine Breakthroughs Changing Treatment Obesity clinical trials are at the forefront of a major transformation in how we understand and treat one of the world’s most pressing public health challenges. Obesity is rising globally at...
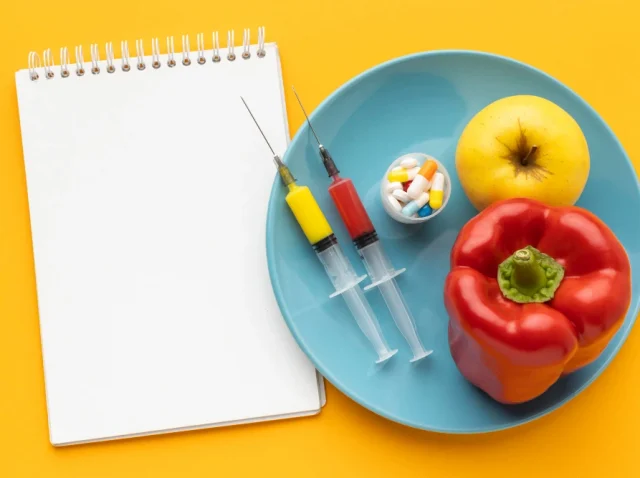
Sustainable Ways to Combat Obesity in Adults Through Lifestyle Changes
Combat Obesity in Adults: 5 Proven Sustainable Lifestyle Changes The most effective way to combat obesity in adults over the long term is not through fad diets or quick fixes — it is through sustained, evidence based lifestyle changes. According to...
Understanding the Genetics of Hypertriglyceridemia: From Common to Rare Cases
Hypertriglyceridemia genetics play a central role in determining why some individuals develop dangerously high triglyceride levels while others with similar lifestyles do not. Triglycerides are fats the body uses for energy storage, packaged in lipoprotein particles like very low density lipoproteins...